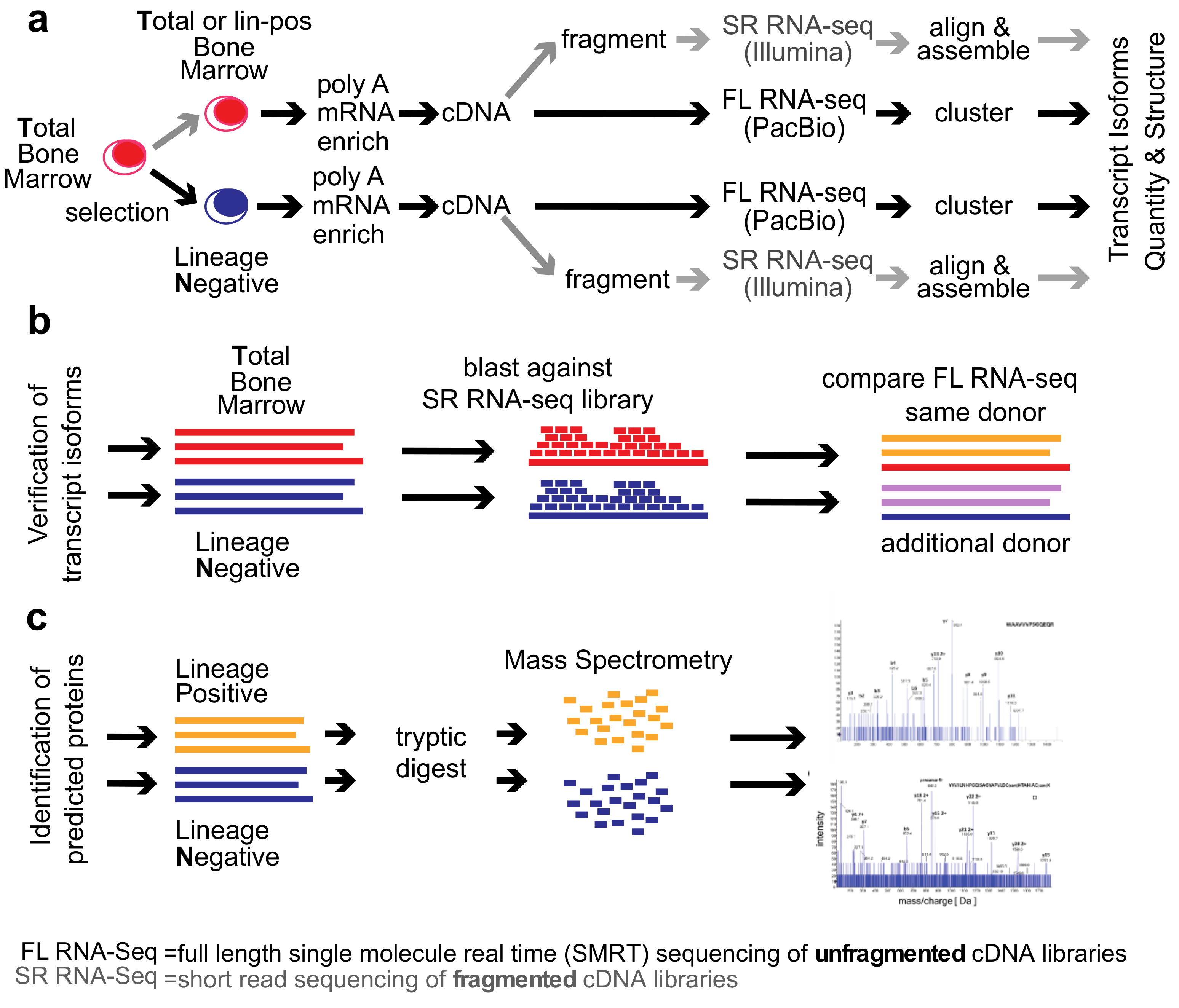
Genes | Free Full-Text | Single-Molecule Real-Time (SMRT) Full-Length RNA-Sequencing Reveals Novel and Distinct mRNA Isoforms in Human Bone Marrow Cell Subpopulations
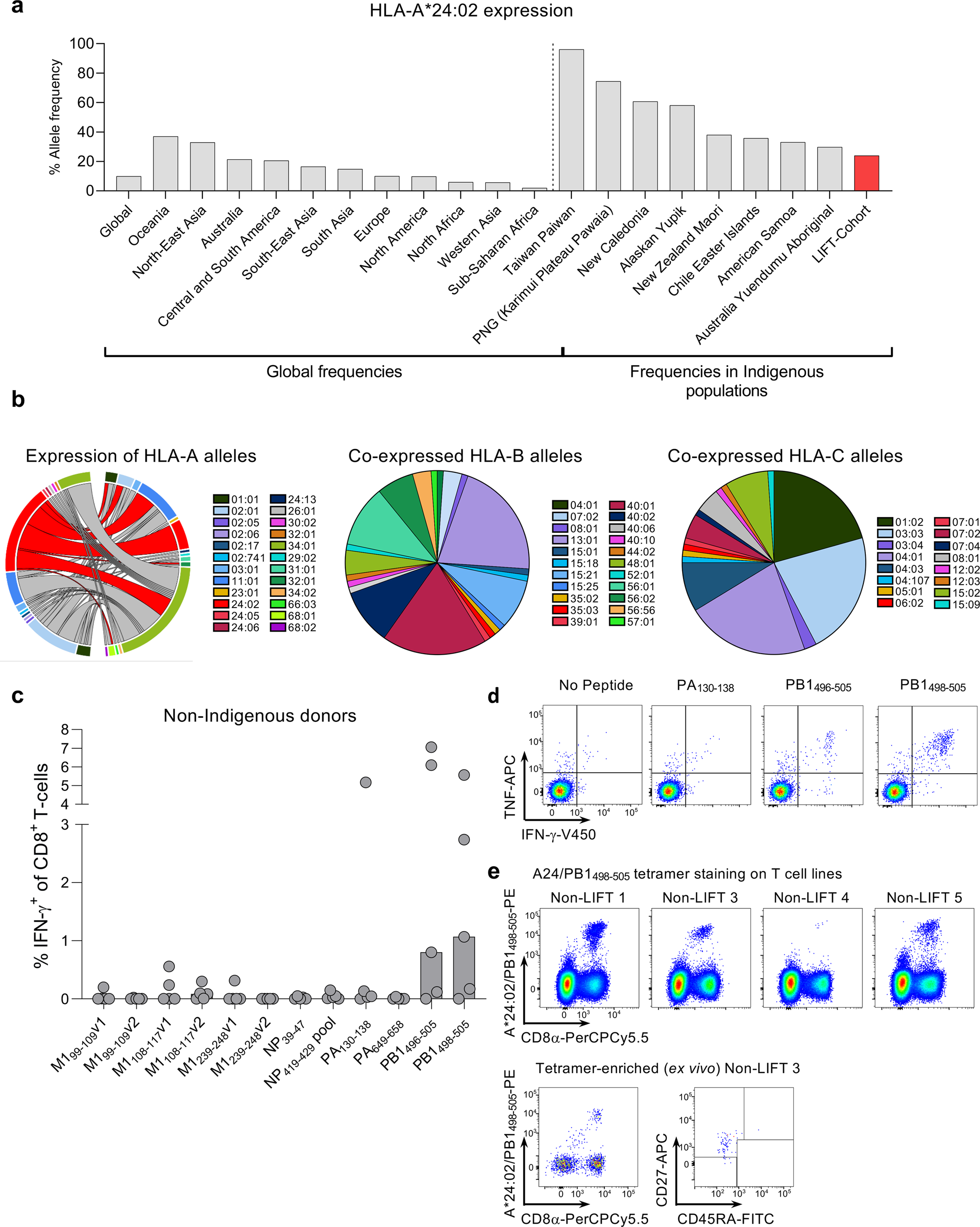
CD8+ T cell landscape in Indigenous and non-Indigenous people restricted by influenza mortality-associated HLA-A*24:02 allomorph | Nature Communications

Repertoire-scale determination of class II MHC peptide binding via yeast display improves antigen prediction | Nature Communications

Evolutionary origins and interactomes of human, young microproteins and small peptides translated from short open reading frames - ScienceDirect

Single-cell analyses reveal key immune cell subsets associated with response to PD-L1 blockade in triple-negative breast cancer - ScienceDirect

Challenges and opportunities associated with rare-variant pharmacogenomics: Trends in Pharmacological Sciences
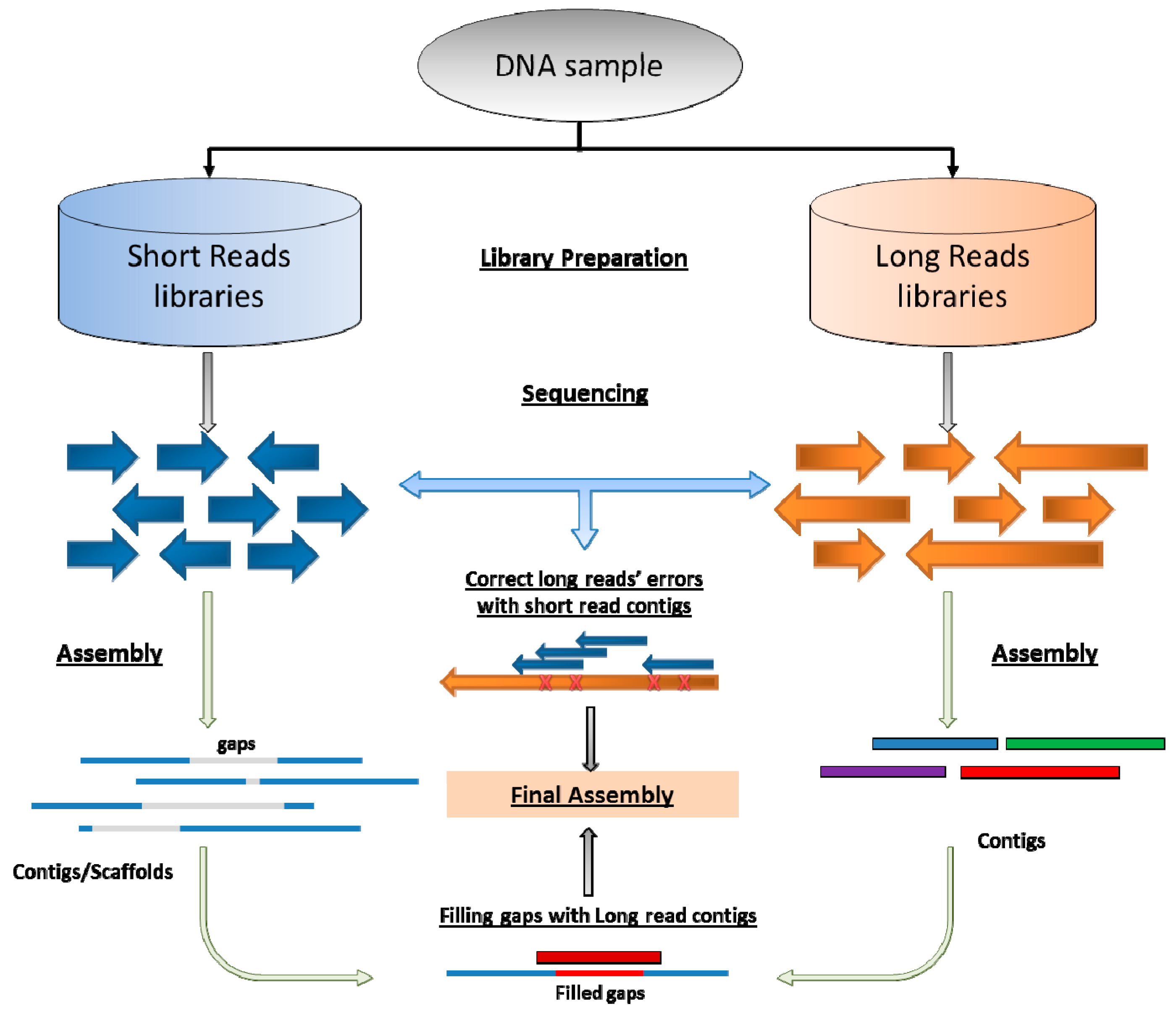
Pharmaceutics | Free Full-Text | Challenges, Solutions, and Quality Metrics of Personal Genome Assembly in Advancing Precision Medicine

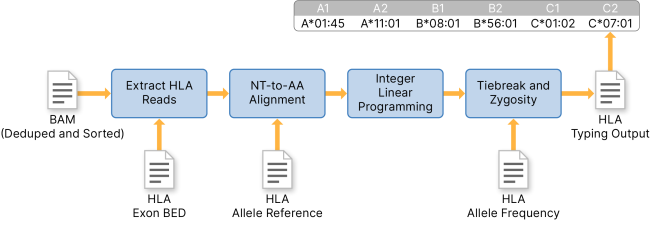
Systematic genetic analysis of the MHC region reveals mechanistic underpinnings of HLA type associations with disease | eLife
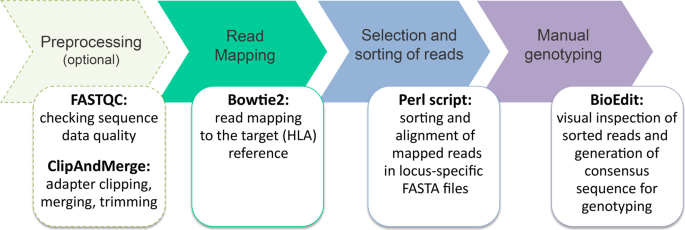
Targeted analysis of polymorphic loci from low-coverage shotgun sequence data allows accurate genotyping of HLA genes in historical human populations | Scientific Reports

Predicting the Affinity of Peptides to Major Histocompatibility Complex Class II by Scoring Molecular Dynamics Simulations | Journal of Chemical Information and Modeling

Generation of hypoimmunogenic T cells from genetically engineered allogeneic human induced pluripotent stem cells | Nature Biomedical Engineering
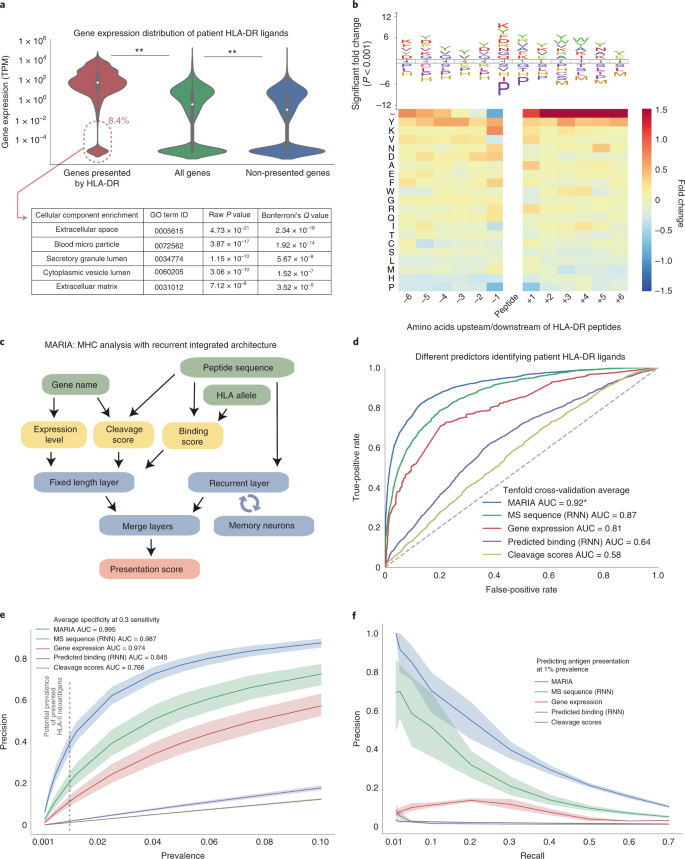
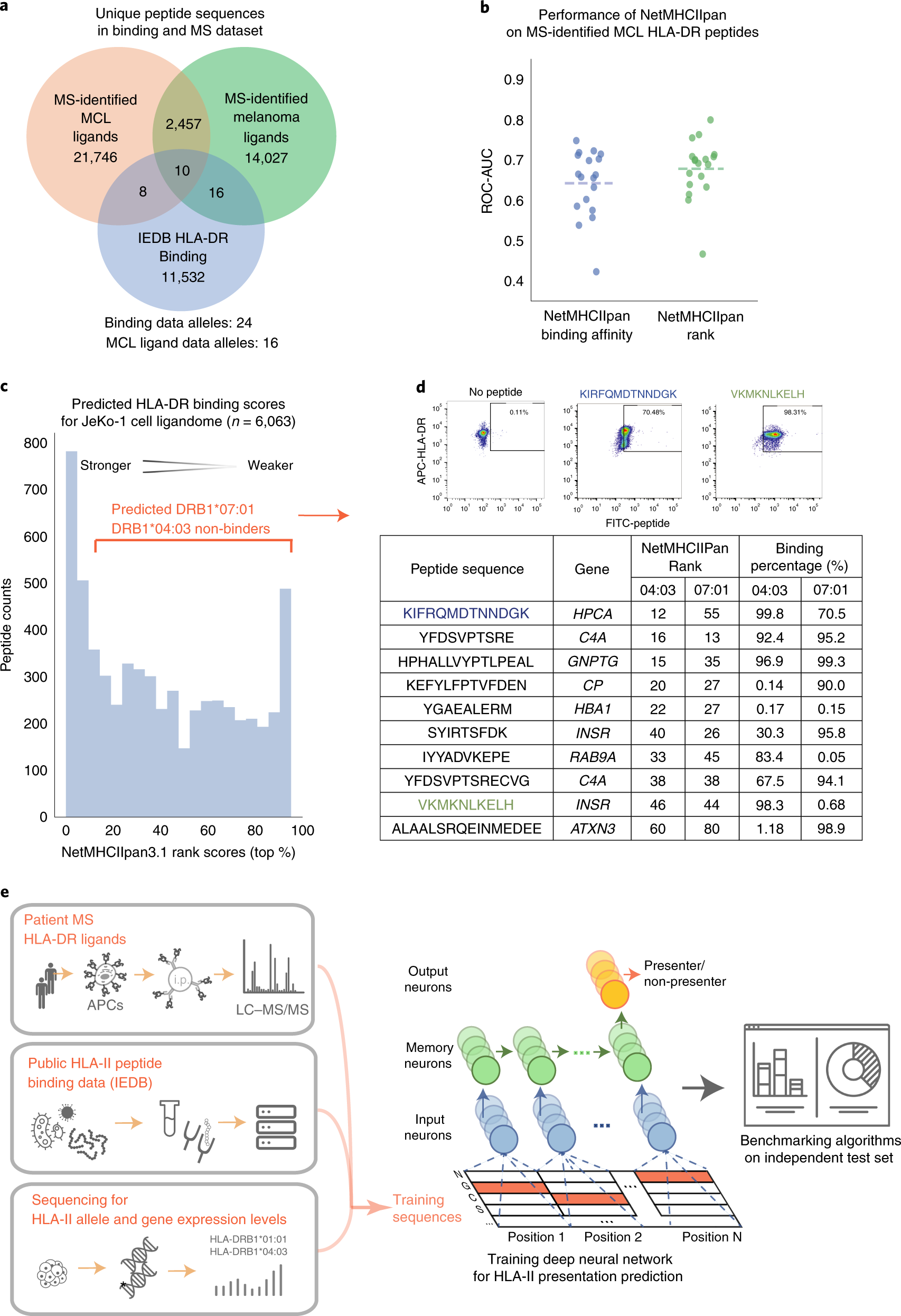
Predicting HLA class II antigen presentation through integrated deep learning | Nature Biotechnology
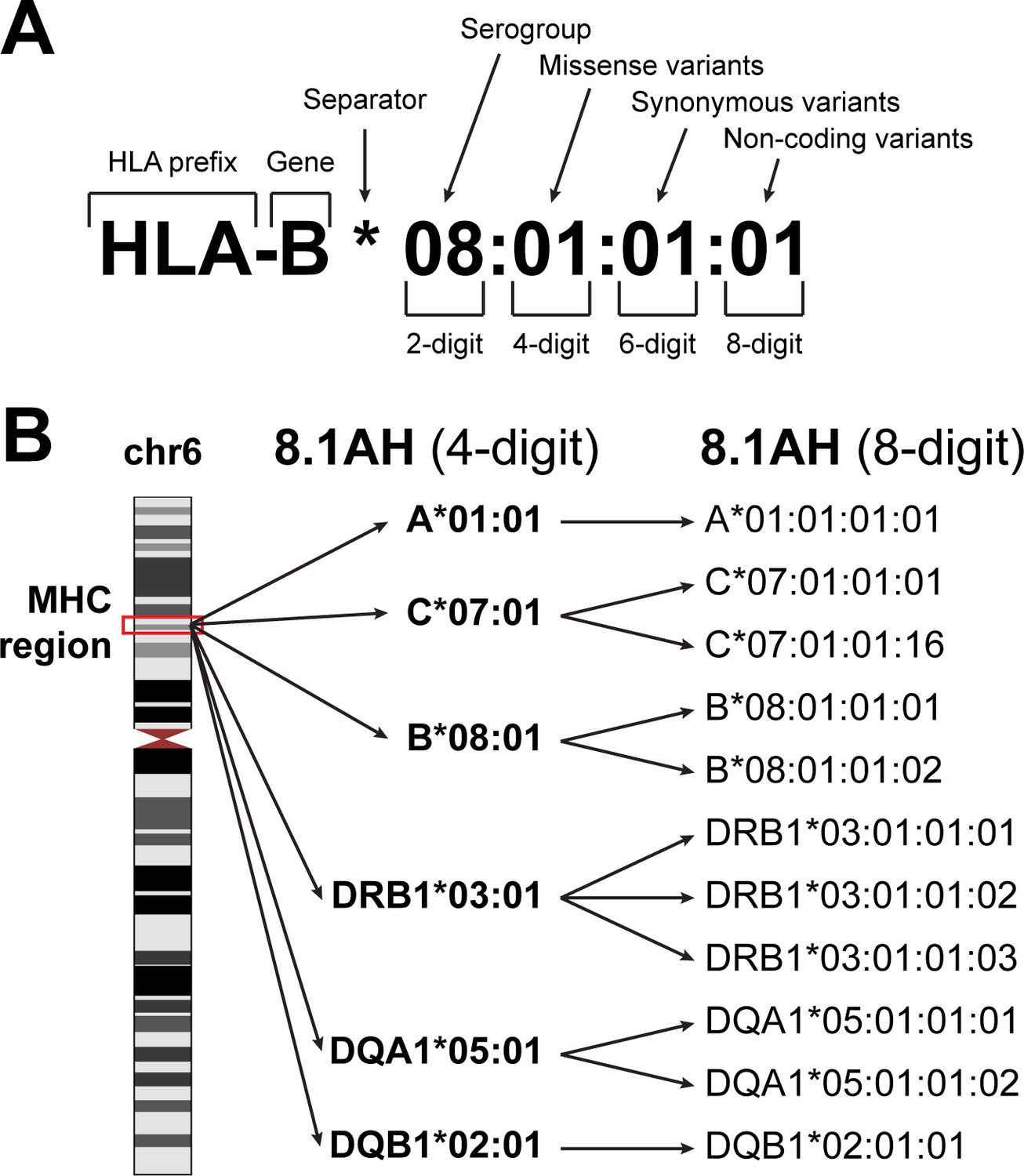
Systematic genetic analysis of the MHC region reveals mechanistic underpinnings of HLA type associations with disease | eLife
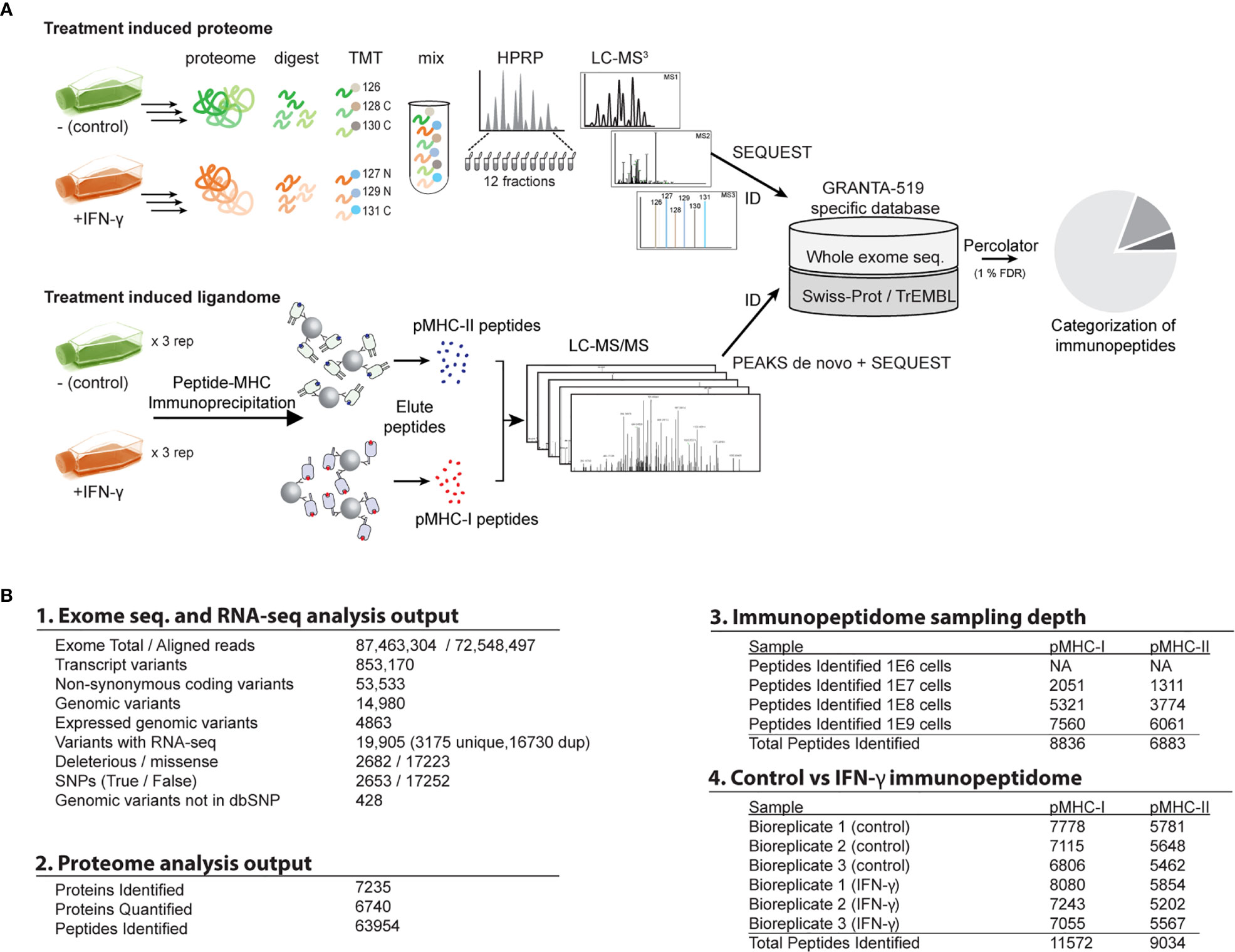
Frontiers | An Integrated Genomic, Proteomic, and Immunopeptidomic Approach to Discover Treatment-Induced Neoantigens
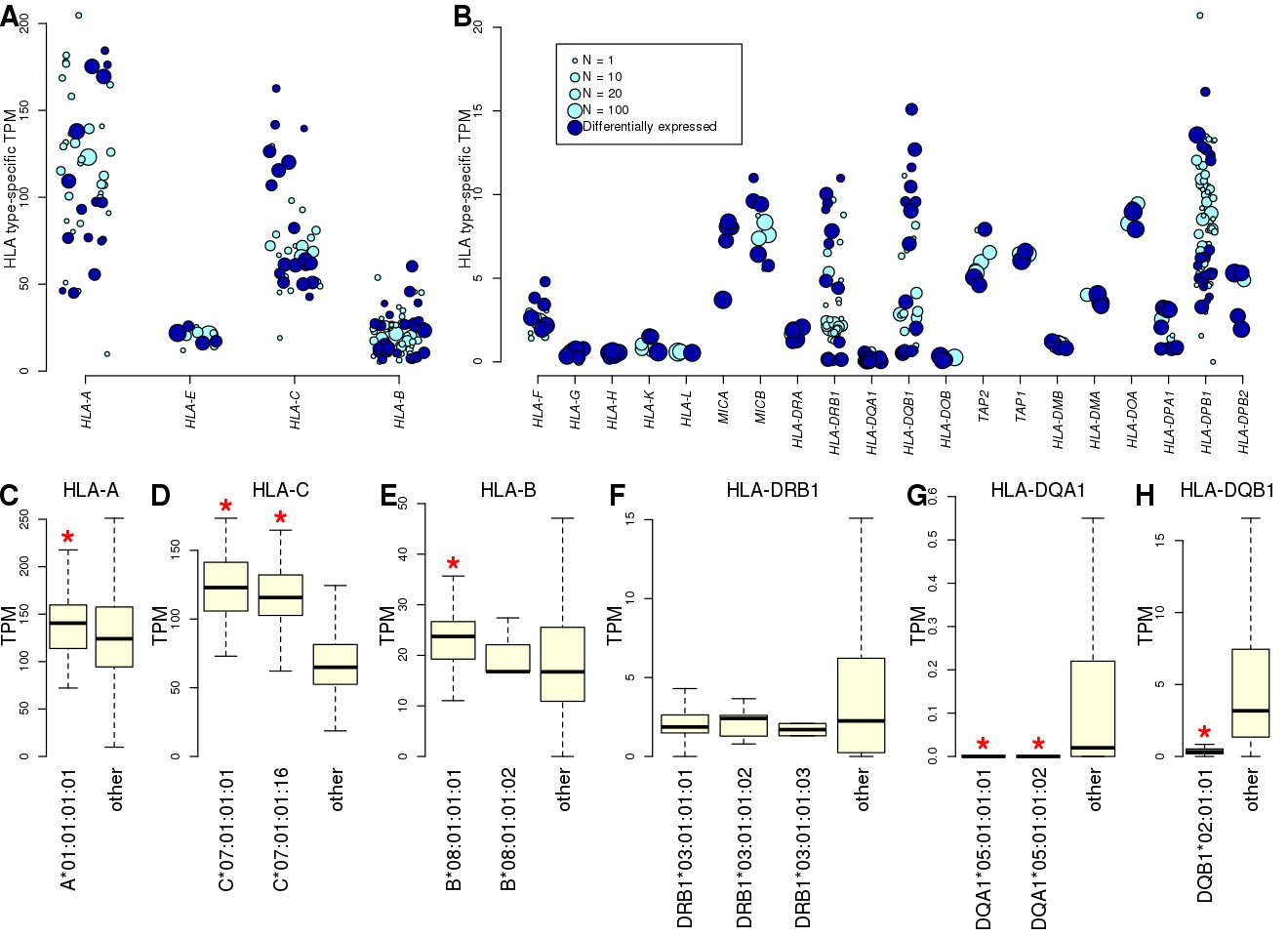
Systematic genetic analysis of the MHC region reveals mechanistic underpinnings of HLA type associations with disease | eLife

Protein-Based Virtual Screening Tools Applied for RNA–Ligand Docking Identify New Binders of the preQ1-Riboswitch | Journal of Chemical Information and Modeling

Identification of neoantigens in oesophageal adenocarcinoma - Nicholas - Immunology - Wiley Online Library

Predicting the Likelihood of Molecules to Act as Modulators of Protein–Protein Interactions | Journal of Chemical Information and Modeling
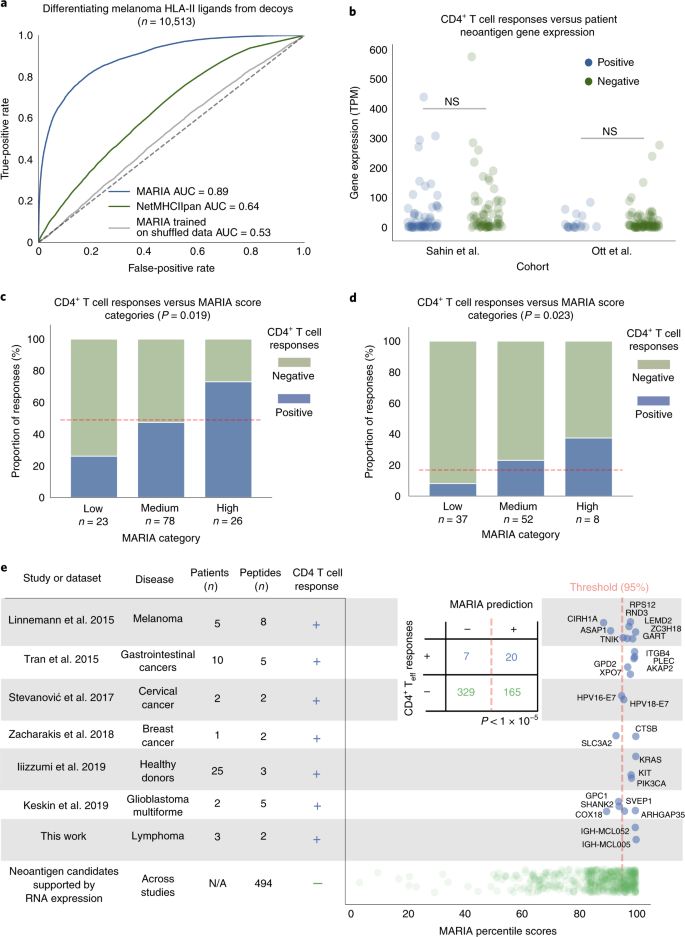
Predicting HLA class II antigen presentation through integrated deep learning | Nature Biotechnology

Deep learning boosts sensitivity of mass spectrometry-based immunopeptidomics | Nature Communications

Proteogenomic Analysis Unveils the HLA Class I-Presented Immunopeptidome in Melanoma and EGFR-Mutant Lung Adenocarcinoma - ScienceDirect